EMASE-Zero is a C++ implementation of EMASE, Expectation-Maximization (EM) algorithm for Allele Specific Expression (ASE). Unlike other approaches, we take into account the nested relationship of isoforms and alleles under genes when we disambiguate multiply aligning reads, which improves downstream analyses performed at the gene level. In combination with alntools (an alignment file processor that generates compressed three-dimensional incidence matrix), Zero estimates the expected read counts fast, over 10 times faster than RSEM. Zero also generalizes our fast hierarchical EM to any decent alignment strategies. The original python implementation is available at https://github.com/churchill-lab/emase.
- Free software: MIT license
Installation
Building and installing EMASE-Zero requires a C++ compiler. After unzipping or extracting the downloaded file, please do the following:
$ cd src
$ make
This will create an executable called emase-zero in the src directory. You can copy this executable into your desired bin directory (such as /usr/local/bin).
Usage
EMASE-Zero takes in an alignment incidence matrix file (a binary format) generated by alntools using the diploid transcriptome alignment BAM file and two text files containing transcript lengths and gene-to-transcript mapping information. We need to specify which EMASE Model to use for disambiguating the origins of multiply-aligning reads.
$ emase-zero -h
EMASE-Zero Version 0.3.1 Help
-----------------------------
USAGE: emase-zero [options]
INPUT: Binary Alignment Incidence file prepared with alntools
(https://churchill-lab.github.io/alntools/)
OPTIONS:
--model (-m) :
Specify normalization model (can be 1-4, default=1)
--output (-o) :
Specify filename for output file (default is input_basename.stacksum.tsv)
--transcript-lengths (-l) :
Filename for transcript length file. Format of each line is
"TranscriptName_HaplotypeName\tlength"
--gene-mappings (-g) :
Filename containing transcript to gene mapping. Tab delimited file where
the first field is the gene ID, all other fields are the transcript IDs
that belong to that gene
--compact-results (-c):
Omit totals that are equal to 0
--no-length-correction (-n):
Do not use transcript lengths as correction
--samples (-s) :
Specify the sample indices. Either one number or in the format
of :
--tolerance (-t) :
Specify the convergence threshold. emase will terminate when the
sum of the aboslute value of differences in the stack sum from one
iteration to the next is lower than this value multiplied by
1,000,000 if performing length adjustment and multipled by #reads if
not (Default = 0.0001)
--max-iterations (-i) :
Specify the maximum number of EM iterations. (Default 999)
--verbose (-v):
Run in verbose mode
--version:
Print the version and exit
--help (-h):
Print this message and exit
•-b or --bin: Specify the name of binary equivalence class (EC) file. A binary EC file is generated from a bam file using alntools. EMASE-Zero does not require any specific aligner to use, but we recommend bowtie Please see example below for the bowtie options we use for RNA-Seq analyses.
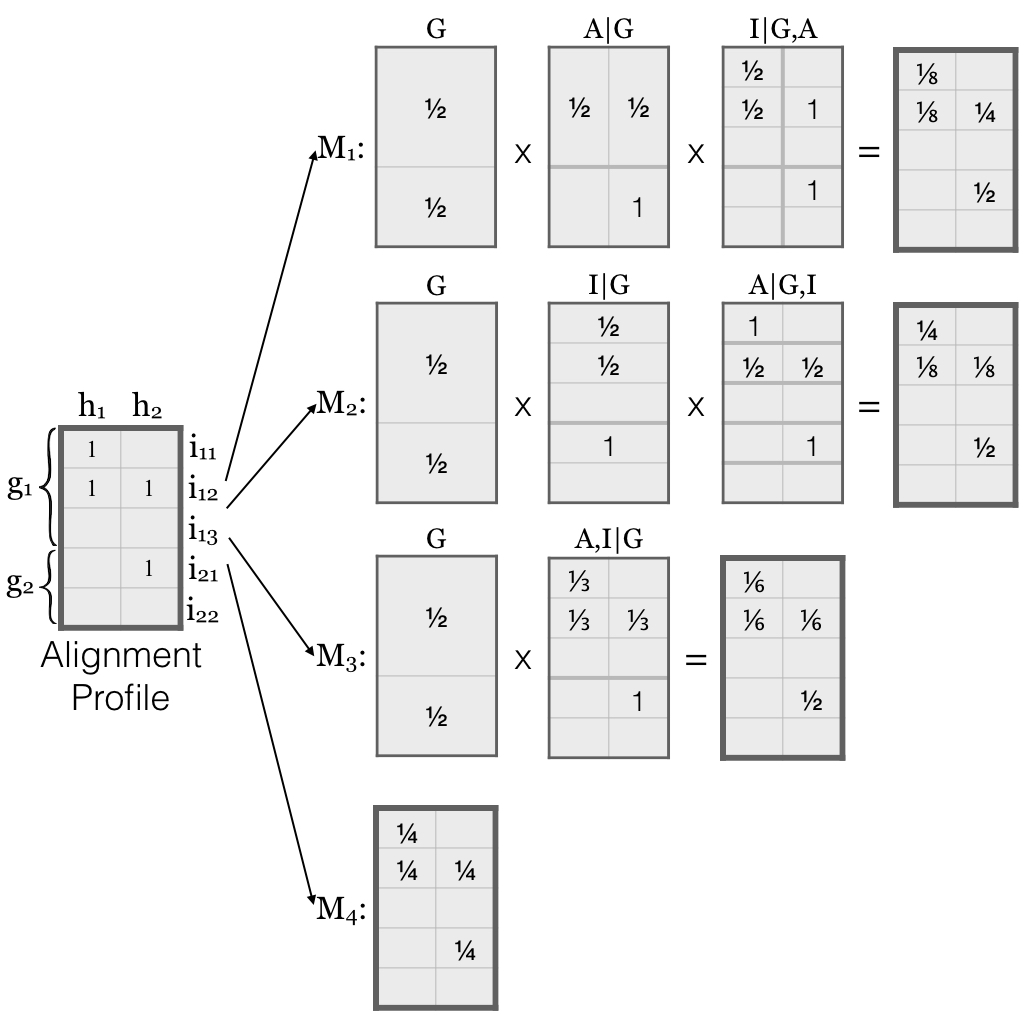
•-m or --model: Specify EMASE model for disambiguating multireads. According to our benchmarking, Model 2 worked best for ASE while Model 4 were the best for total gene exression.
•-t or --transcript-lengths: Specify transcript length file name. We assume the file has the format of "Transcript_ID[tab]Length". For example, assuming 'M' and 'P' stand for maternal and paternal alleles respectively,
ENSMUST00000000001_M 3262
...
ENSMUST00000098609_M 2653
...
ENSMUST00000000001_P 3264
...
ENSMUST00000098609_P 2659
...
•-g or --gene-mappings: Specify gene-to-transcripts mapping file name. The file should be formatted in Gene_ID[tab]Transcript_ID_1[tab]Transcript_ID_2[tab]…. These mappings are among reference IDs, not among allele IDs, so no haplotype code at the end like _M or _P. For example,
ENSMUSG00000000001 ENSMUST00000000001
ENSMUSG00000000003 ENSMUST00000000003 ENSMUST00000114041
ENSMUSG00000000028 ENSMUST00000000028 ENSMUST00000096990 ENSMUST00000115585
...
Here's an example pipeline of commands:
$ bowtie -q -a --best --strata --sam -v 3 -m 100 bowtie.transcriptome my_sample.fastq | samtools view -bS - > my_sample.bam
$ alntools bam2ec -t transcripts.lengths.tsv -c 8 my_sample.bam my_sample.bin
$ emase-zero -m 2 -b my_sample.bin -o emase-zero.quantified -l transcript.lengths.tsv -g gene2transcripts.tsv
Note: We highly recommend adding poly-A tail to the alignment target sequences (transcripts) when you build bowtie index. We usually add (R-1)-long poly-A tails where R is the read length.
Announcements
- 12 Feb 2018 » How to cite this work
- 06 Apr 2017 » Use alntools to make binary EC file
- 26 Jun 2015 » Welcome to EMASE-Zero!